Omics Beyond Transcriptomics Session Summary
Denina Simmons, Ontario Tech University; Ksenia Groh, EAWAG
At the SETAC Europe meeting in Seville, we had the pleasure of co-chairing the session, “Omics Beyond Transcriptomics: Leveraging Proteomics and Metabolomics to Improve Mechanistic Understanding of Responses to Environmental Stressors.”
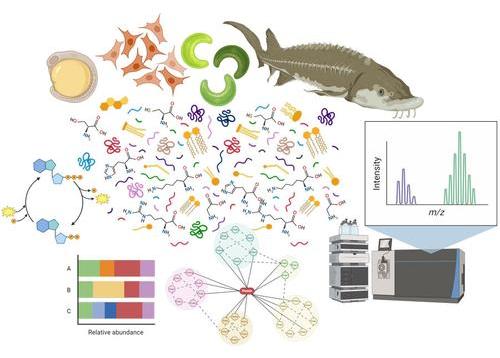
While there are many varying definitions of “omics,” our favorite descriptive analogy is that a single invoice is to economics what a single molecule is to omics. Imagine you’re the mayor of a vibrant city. Since you can’t manage the city’s budget by looking at one invoice at a time, you would likely invest in an advanced city management system. This system would provide you with a complete financial overview of every income and expense, from taxes to large infrastructure projects. In good and bad times, you will be able to quickly assess how the different parts of the city interact and find the right balance for the competing needs. Similarly, omics lets you see how multiple biomolecules interact in a biological system, so you can understand their interdependencies and their collective responses to any internal or external perturbations.
In environmental toxicology, omics tools can be used to measure many molecular endpoints all at once, providing a more complete and unbiased view of biological responses, which could allow us to understand the toxicological mechanisms and impacts of chemicals and other stressors on living organisms. However, among all the different possibilities of looking at what’s out there in the cells and organisms, transcriptomics has become the most common type of omics tools used by biologists working in environmental toxicology so far. This is likely because the transition from qPCR to sequencers was more intuitive and seamless, while the measurement of proteins and metabolites requires completely different instrumentation (often chromatography coupled with mass-spectrometry), which is expensive and could require a greater learning curve for new users to master. Yet, because proteins and metabolites are the end-products of genes and transcripts, and as such bear the closest links to (altered) phenotypes, it is equally, if not more, important to measure these biomolecules in order to understand their role in toxicological responses. Coming back to our financial analogy, limiting the omics analysis to transcriptomics only would be like trying to develop a budget by accounting only for transportation and school education costs, while neglecting the needs of police corps and health care services.
The goal of our session was thus to spotlight the recent technical advancements and the rising potential of the “other omics,” which include proteomics, lipidomics and metabolomics among others, to increase our ability to assess the toxicological effects of chemicals, compared with the use of transcriptomics alone. With 12 and 20 abstracts submitted for platform and poster presentations respectively, we faced a tough challenge, because our short session slot had room for only five platform talks. We therefore selected only three abstracts for full platform talks and used the remaining session time to spotlight six additional studies from the remaining 29 abstracts. This allowed us to feature nine research works instead of only five. The session began with presentations on metabolomics and proceeded to proteomics and multi-omics studies. Overall, the platform session was a success and the poster session was packed as well! Some of the posters included interactive aspects, with flaps, games and fun sticker handouts, which greatly enhanced interactions with the audience.
The breadth of investigated stressors ranged from individual substances like 6-phenyl-1,3,5-triazine-2,4-diamine-4-oxide quinone (6PPD quinone), pesticides, antibiotics, phthalates, pharmaceuticals, and per- and polyfluoroalkyl substances (PFAS) to microplastics, as well as chemical mixtures and cumulative stressor impacts. Beyond the transcriptome, the studies focused on the metabolome, lipidome and (phospho)proteome, as well as the phenome (i.e., the complete set of phenotypes expressed by an organism) of different specimens, including periphyton, plants, corals, invertebrates, endangered fish species and alternative test models such as human and animal cell cultures, as well as zebrafish embryos. Collectively, these studies highlighted the applications and usefulness of different types of omics approaches to assess chemical effects, decipher mechanisms of toxicity and identify biomarkers of exposure and effects that can be used to support both the prospective risk assessment and the environmental biomonitoring needs.
Next year, we would like to host another session with the same goal of showcasing the unique insights that can be delivered by different omics tools. This time, we aspire to challenge all our presenters to really “tell a story” – a biological story of life encountering and counteracting chemical toxicity stress – with their data, as opposed to just showing complex graphs, shiny heatmaps and model estimates. We are also open to incorporating interactive presentation formats, and we would welcome any other innovative ideas that could help to further spread the word about the “other omics,” and their value in human and environmental toxicology. If you are currently working in the omics space, and if you are including the “other omics” tools in your work, please consider submitting an abstract to our session in Vienna at SETAC Europe 2025!
Authors’ contact: [email protected], [email protected]